Getting Started
ParamRF provides a declarative modelling interface that compiles RF models, such as circuit models, using JAX. This page provides in introduction into how to define models, as well as fit them.
Core Concepts
The library revolves around a few key building blocks:
pmrf.Model: The base class for any RF model. When inherited from, methods such as s, a, z and y can be overriden to define model S-parameters, ABCD-parameters etc. These methods all accept frequency as input. On the other hand, __call__ can be overridden to return a model instance itself, for more complex compositional model building.pmrf.Frequency: A wrapper around a JAX array that defines the frequency axis over which models are evaluated.parax.Parameter: A parameter in a model (from parax), storing its value and metadata. This allows for parameter bounds and scaling, marking parameters as fixed, and associating a distribution with the parameter for Bayesian fitting.
pmrf.Evaluator: A lower level object that is used to extract features from a model in a composable manner. These are created under-the-hood e.g. when specifying custom features for fitting, but can also be specified directly for advanced cost and likelihood functions.
Model Composition
ParamRF provides a component library with commonly-used models such as lumped and distributed elements. Models can be built directly using these in a compositional approach.
Cascaded Models
For simple circuit element chains, the ** operator can be used to cascade several models together.
The example below creates an RLC filter and terminates it in an open circuit. The resultant rlc is a first-class pmrf.Model of type pmrf.models.Cascade, consisting of parameters representing the respective R, L and C parameters. The S11 is then plotted using scikit-rf under the hood.
from parax import Parameter, Fixed
import pmrf as prf
from pmrf.models import Resistor, Inductor, ShuntCapacitor, OPEN
# Instantiate the lumped element models
resistor = Resistor(R=100.0)
inductor = Inductor(L=Parameter(2.0, scale=1e-9)) # we can optionally provide a parameter scale
capacitor = ShuntCapacitor(C=1.0e-12, name="cap") # naming makes parameter manipulation later easy
# Cascade the models, storing the result.
# We also create a terminated version with a new, fixed C
rlc = resistor ** inductor ** capacitor
terminated_rlc = rlc.terminated(OPEN).with_params(cap_C=Fixed(0.5e-12))
# Plot the S11 of the terminated model at a specified frequency range
freq = prf.Frequency(1, 1000, 1000, 'MHz')
terminated_rlc.plot_s_db(freq, m=0, n=0)
Circuit Models
For complex circuits, ParamRF offers the ability to combine models in any desired configuration using the pmrf.models.Circuit class. This class accepts a list of “connections”. Each entry in this list is a node in the circuit. Each node is another list, with each element being a tuple for each connected circuit element or sub-model. Each tuple then contains the model object, as well as the index of the port for that model that is connected in that node.
The following example uses this method to define a two-port PI-CLC network. “External” nodes (each entry in the outer list) are numbered as E0, E1 etc. whereas “internal” port indices (ports for each model in the circuit) are numbered per element as I0, I1 etc. The model is then converted to a scikit-rf network and plotted.
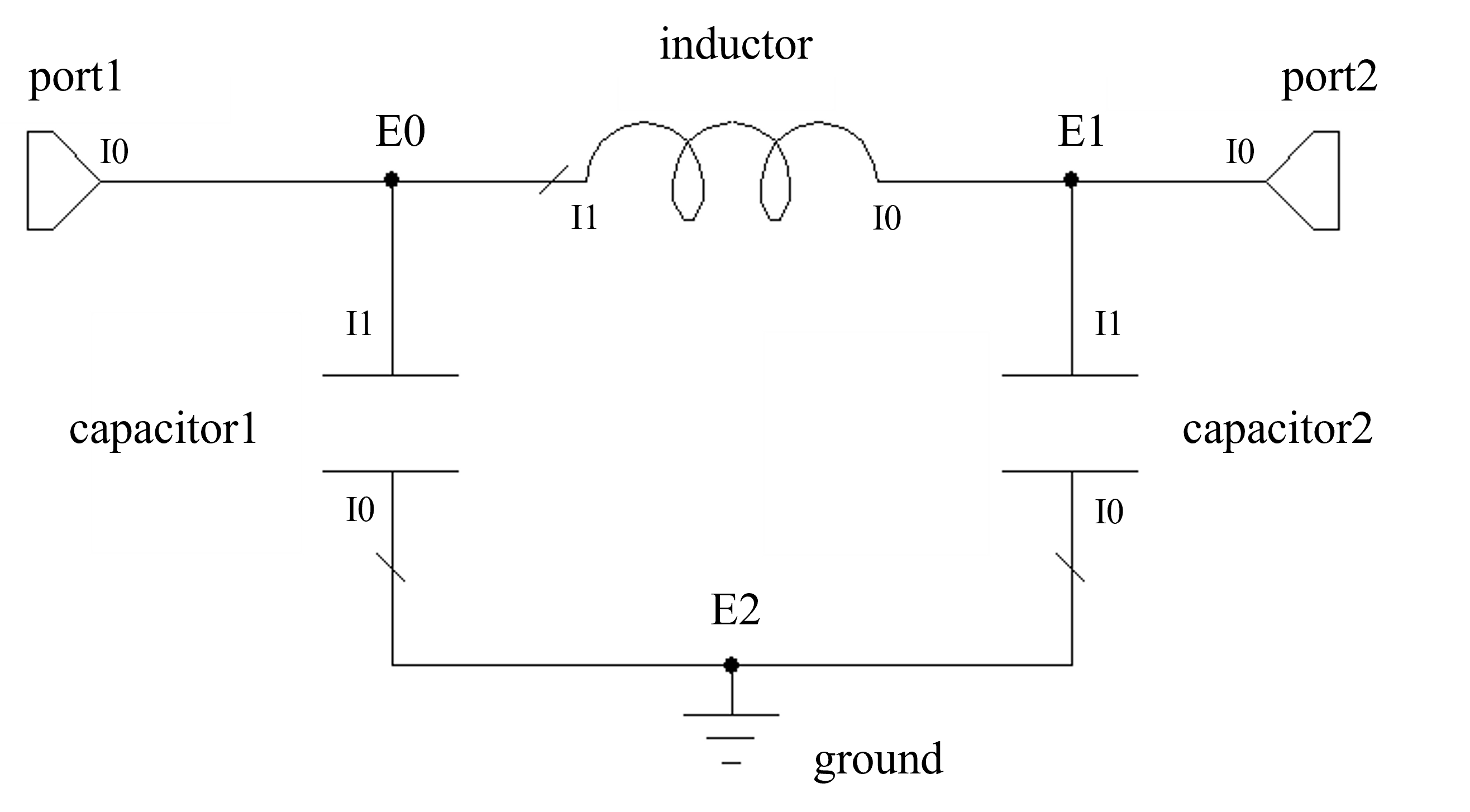
import pmrf as prf
from pmrf.models import Capacitor, Inductor, Circuit, Port, Ground
# Instantiate the elements, ports and grounds
capacitor1, capacitor2 = Capacitor(C=2e-12), Capacitor(C=1.5e-12)
inductor = Inductor(L=3e-9)
port1, port2 = Port(), Port()
ground = Ground()
# Create the connections list
connections = [
[(port1, 0), (capacitor1, 1), (inductor, 1)], # E0
[(port2, 0), (capacitor2, 1), (inductor, 0)], # E1
[(ground, 0), (capacitor1, 0), (capacitor2, 0)], # E2
]
# Create the model and plot it's S21 parameter
pi_clc = Circuit(connections)
freq = prf.Frequency(1, 1000, 1001, 'MHz')
pi_clc.plot_s_db(freq, m=1, n=0)
# Note that ParamRF already provides a built in, more efficient PiCLC model
from pmrf.models import PiCLC
PiCLC(2e-12, 3e-9, 1.5e-12).plot_s_db(freq, m=1, n=0)
Model Inheritance
For more complex models (such as equation-based ones), users can inherit directly from the pmrf.Model class and override one of the network properties (such as s, a, or y) or the __call__ method.
Any attributes of a model are classified as either static or dynamic. By default, fields of built-in types such as str, int, list etc. are seen as static in the model hierarchy, whereas those annotated as a parax.Parameter or pmrf.Model are dynamic and can be adjusted (for example, by fitting routines).
Note that parameter initialization is flexible: parameters may be populated with a simple float value; using factory methods such as parax.Uniform, parax..Normal or parax..Fixed; or directly using the parax.Parameter class constructor.
Equation-based Models
The following example demonstrates custom model definition by defining a capacitor from first principles. This could be used, for example, to implement more complex analytic or surrogate models. Here, one of the typical network properties, such as pmrf.Model.s(), pmrf.Model.a(), pmrf.Model.y(), or pmrf.Model.z(), must be overriden, returning the resultant matrix directly.
import jax.numpy as jnp
import pmrf as prf
from parax import Parameter
# Define a model class. Behaviour is defined by implementing
# a primary matrix function such as "s" in this case.
class Capacitor(prf.Model):
C: Parameter = 1.0e-12
def s(self, freq: prf.Frequency) -> jnp.ndarray:
w = freq.w
C = self.C
z0_0 = z0_1 = self.z0
denom = 1.0 + 1j * w * C * (z0_0 + z0_1)
s11 = (1.0 - 1j * w * C * (jnp.conj(z0_0) - z0_1) ) / denom
s22 = (1.0 - 1j * w * C * (jnp.conj(z0_1) - z0_0) ) / denom
s12 = s21 = (2j * w * C * (z0_0.real * z0_1.real)**0.5) / denom
return jnp.array([
[s11, s12],
[s21, s22]
]).transpose(2, 0, 1)
Circuit Models
Sometimes it is still convenient to inherit from pmrf.Model while still building the model using cascading or pmrf.models.Circuit. In this case, the model can be built from sub-models fields/attributes, and returned by overriding the pmrf.Model.__call__() method.
The following example creates a PI-CLC model once again, but using the above method. Note how certain parameters can be given initial parameters, bounds or fixed to a constant (useful for fitting).
from parax.parameters import Uniform, Fixed
import pmrf as prf
from pmrf.models import Capacitor, Inductor, Circuit, Port, Ground
class PiCLC(prf.Model):
capacitor1: Capacitor = Capacitor(C=Fixed(1.0e-12))
capacitor2: Capacitor = Capacitor(C=Uniform(0.0, 10.0, value=2.0, scale=1e-12))
inductor: Inductor = Inductor(L=Uniform(0.0, 10.0, value=2.0, scale=1e-12))
def __call__(self) -> prf.Model:
# Instantiate the ports and grounds
port1, port2, ground = Port(), Port(), Ground()
# Create the connections list. This time, capacitor1, capacitor2 and inductor are members.
connections = [
[(port1, 0), (self.capacitor1, 1), (self.inductor, 1)], # E0
[(port2, 0), (self.capacitor2, 1), (self.inductor, 0)], # E1
[(ground, 0), (self.capacitor1, 0), (self.capacitor2, 0)], # E2
]
# Return the model
return Circuit(connections)
Fitting
Models can easily be fit using to measured data using the pmrf.fit module. The general workflow consists of defining a model, loading data via scikit-rf, initializing the solver (optimizer/inferer), defining the features to optimize (e.g. S11), and running the fit.
Solvers
ParamRF allows for optimization using either scipy.optimize.minimize or optimistix.minimise, and Bayesian inference using inferix, which provides wrappers for PolyChord and BlackJAX.
Scipy: Provides a wrapper around gradient-based and gradient-free optimization algorithms from
scipy.optimizeinpmrf.optimize.ScipyMinimizer. This includes algorithms such as SLSQP, Nelder-Mead and L-BFGS. These algorithms are CPU-native and cannot run on the GPU.Optimistix: Provides JAX-native optimization algorithms, such as
optimistix.BFGSandoptimistix.NelderMead. These algorithms run their loop directly in JAX, and therefore can be compiled to any architecture (CPU, GPU, TPU).Inferix: Enables Bayesian inference through nested sampling and MCMC sampling using e.g.
inferix.PolyChordandinferix.NUTS. This approach provides maximum likelihood parameters, as well as full posterior probability distributions and Bayesian evidence for model comparison. We recommend this source for a brief introduction to nested sampling and Bayesian inference.
Example
The following provides an example of fitting the built in pmrf.models.CoaxialLine model to the measurement of 10m coaxial cable (provided as an example in the GitHub). Data is loaded using scikit-rf, the model is instantiated with appropriate initial parameters, the fit is run, and results are plotted.
import logging
import skrf as rf
from parax.parameters import Uniform, RelativeNormal, Fixed
from pmrf.models import CoaxialLine
from pmrf.optimize import fit, ScipyMinimizer
logging.basicConfig(level=logging.INFO)
# Load the measured data.
measured = rf.Network('data/10m_cable.s2p', f_unit='MHz')
# Setup the model. Note that any parameters not passed
# are set as free with infinite bounds
model = CoaxialLine(
din = RelativeNormal(1.12, 0.05, scale=1e-3),
dout = RelativeNormal(3.2, 0.05, scale=1e-3),
epr = Fixed(1.384),
rho = RelativeNormal(1.6, 0.05, scale=1e-8),
tand = Uniform(0.0, 0.01, value=0.0, scale=0.01),
length = RelativeNormal(10.0, 0.05),
mur = Fixed(1.0),
)
# Fit the model
results = fit(model, measured, solver=ScipyMinimizer())
fitted_model = results.model
# Plot some results
fitted_model.to_skrf(measured.frequency).plot_s_db()
measured.plot_s_db()